ICPL™ Kit by SERVA Electrophoresis
A powerful technology for comparative quantification of independent proteins.
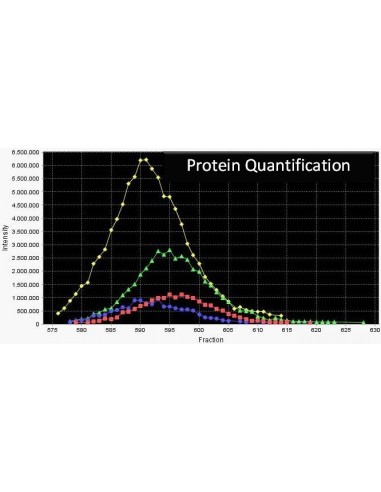
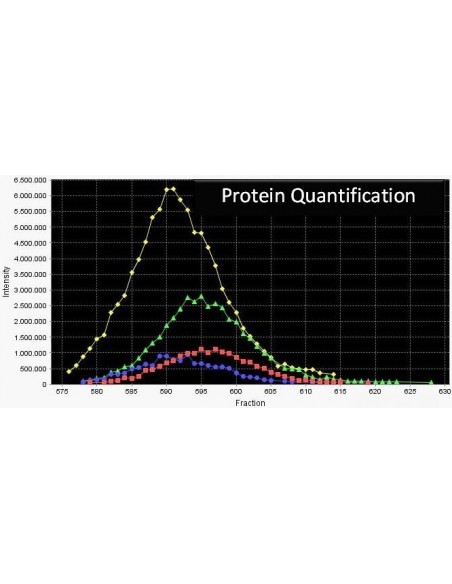
Serva ICPL ™ is based on the powerful technology of isotope coded protein labeling (ICPL) of all lysine residues and N-termini of intact proteins; the labeling step is done at the protein level and includes all free amino acid groups. In contrast, with alternative methods the labeling step is only performed post proteolytic digestion, that is on the peptide level. Therefore the analytical depth of a proteome analysis is improved significantly. For the stable labeling of intact proteins nicotinic acid-NHS-12C respectively nicotinic acid-NHS-13C is used.
The kit contains reagents for 2 x 6 reactions:
- Two Labels: 12C- and 13C-Nic-reagent
- Stop solution 1 + 2
- Reduction solution
- Alkylation reagent
- Lysis buffer
- Standard protein mix A + B
This technology is applicable to any protein samples, including body fluids and tissue extracts. It reduces the proteome complexity by fractionation of intact proteins thus enabling high-throughput quantitative protein profiling. ICPL™ method allows users to analyse post-translational modifications, processing variants, membrane proteins as well as protein isoforms and is compatible with all separation methods currently employed in proteome studies.
Freeware… ICPLQuant software enables the user to quantify and identify even low abundance proteins in a complex mixture simultaneously. ICPLQuant takes deisotoped data as input, generates precursor files for MS/MS and is able to import MASCOT result files for combining MS/MS and quantification results. By selecting only regulated precursors for subsequent MS/MS experiments, the software minimizes time consuming acquisition and interpretation of MS/MS data. ICPLQuant is based in two main and independent units. Unit 1 is designated for ICPL multiplex quantification and unit 2 for ICPL multiplex identification. The second unit interacts with MASCOT and parses MSMS. The software tool was developed by TopLab GmbH based in Martinsried Germany by Lottspeich et al. and is marketed worldwide through SERVA. ICPLQuant is free of charge at http://www.biochem.mpg.de/en/rg/lottspeich/technologies/ICPLQuant/index.html